Our Services
Zetobit serves clients in the biotech, pharma, and life sciences industry — and the academic groups that collaborate with them — delivering bioinformatics expertise across the full research pipeline. Whether you're accelerating drug discovery, building a genomics platform, or advancing a research program, we curate datasets, mine public repositories, design statistically rigorous studies, run state-of-the-art sequencing, and deliver deep analysis with compelling visualizations. Already mid-project? We plug in exactly where you need us.
Institutional-grade bioinformatics. Built for biotech, pharma, and the research teams that drive them. From data curation to publication-ready results — we plug in wherever you need us. Book a free consultation →
Curation
Analysis
Design
Services
& Visualization
in here
in here
in here
in here
in here
Data Curation
Reliable analysis begins with reliable data. We specialize in end-to-end data curation — sourcing, cleaning, harmonizing, and annotating datasets from diverse origins including proprietary experimental results, electronic health records, and major public repositories such as GEO, TCGA, UK Biobank, dbGaP, and CCLE. Our team resolves inconsistencies in metadata, standardizes nomenclature across platforms and cohorts, flags and removes low-quality samples, and structures data into analysis-ready formats. The result is a curated, reproducible dataset that your team can trust — and that holds up to regulatory and peer scrutiny alike.
Starting with messy or inconsistent data? That's exactly where we begin. Tell us about your dataset and we'll outline how we'd approach it. Schedule a scoping call →




Public Dataset Analysis
The world's leading biological databases hold answers your next experiment hasn't asked yet. We mine and reanalyze data from repositories including GEO, SRA, TCGA, UK Biobank, ENCODE, dbGaP, and CCLE to surface insights that reduce experimental costs, validate hypotheses before committing resources, and identify biomarkers, expression signatures, and population-level trends hidden in existing data. Our team builds custom pipelines tailored to each repository's data structure and quirks — normalizing across studies, correcting for batch effects, and integrating multi-omics layers to give you a richer, more complete picture. The result is faster, evidence-backed decision-making without the cost and timeline of generating new data from scratch.
Not sure if the data you need already exists? It probably does. We'll help you find it, reanalyze it, and extract what's relevant to your program — before you spend on new experiments. Talk to our team →


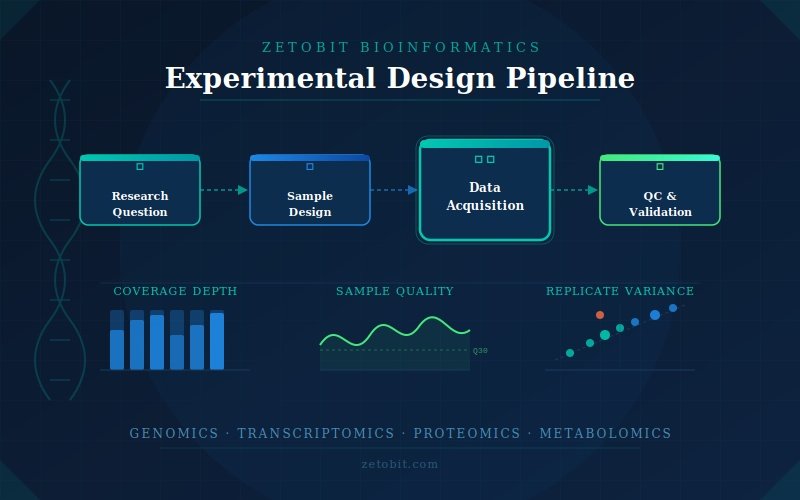
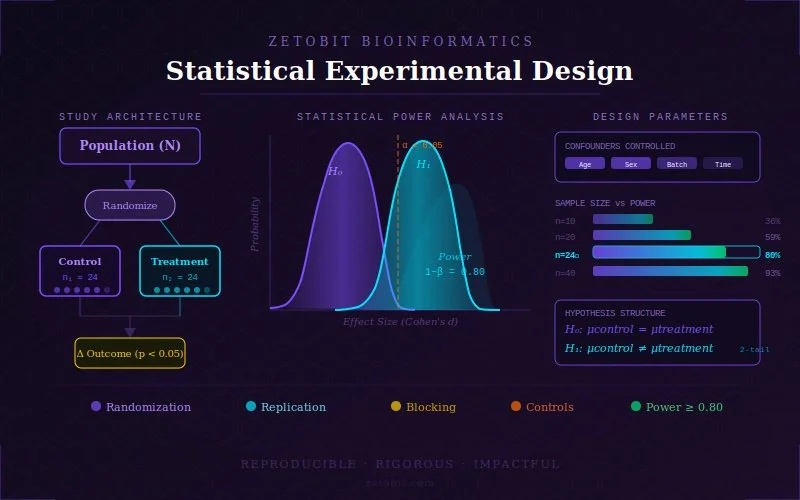
Experimental Design
A poorly designed experiment cannot be rescued by even the most sophisticated analysis. Our experimental design consultations are built around your specific biological question, sample constraints, and downstream analytical goals. We advise on study architecture — including case-control frameworks, longitudinal designs, and multi-cohort strategies — and determine the statistical power needed to detect meaningful effect sizes with confidence. We guide sequencing platform and assay selection, recommend appropriate sample sizes, define inclusion and exclusion criteria, and establish quality thresholds before a single sample is processed. For multi-omics studies, we design integrated collection strategies that ensure each data layer is compatible and analytically coherent. The goal is a study that generates clean, high-confidence data from day one — and stands up to the scrutiny of both internal review and external publication.
Designing a study and want a second set of expert eyes before you commit resources? A single consultation can prevent months of rework. Let's look at your design together. Book a design review →
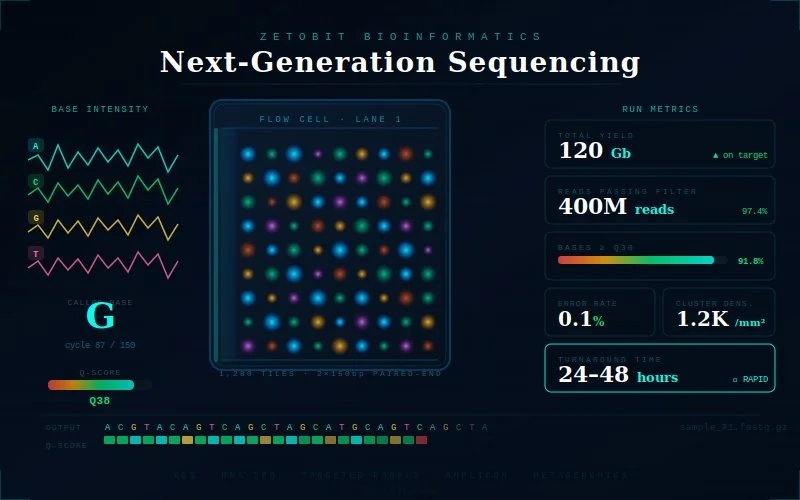
Sequencing Services
We support a comprehensive range of next-generation sequencing applications — including whole genome sequencing (WGS), whole exome sequencing (WES), bulk RNA-seq, single-cell RNA-seq (scRNA-seq), ChIP-seq, ATAC-seq, and targeted panel sequencing — across human, model organism, and non-model species. Every sequencing project is preceded by rigorous library quality assessment and sample QC to ensure data integrity before sequencing begins. Raw data is processed through validated primary analysis workflows — alignment, variant calling, quantification — and delivered in analysis-ready formats alongside comprehensive QC reports. Whether you are sequencing a handful of discovery samples or scaling to a large clinical or population cohort, our workflows are designed to maintain consistency, reproducibility, and high data quality at every scale.
Have samples ready to sequence — or data already in hand? We support projects at any stage, from library prep through final variant report, with rigorous quality standards throughout. Get a project estimate →
Proprietary Dataset Analysis & Visualization
Your proprietary data contains answers that generic pipelines will miss. We deliver end-to-end analysis of your internal datasets — whether derived from sequencing, proteomics, metabolomics, or multi-omics experiments — using analytical frameworks tailored to your specific biological question and program stage. Our team performs differential expression analysis, variant annotation and interpretation, pathway and gene ontology enrichment, survival analysis, dimensionality reduction, and single-cell deconvolution, among other methods. Where relevant, we integrate your proprietary data with curated public datasets to provide comparative context, benchmark your findings against existing literature, and surface novel associations that internal data alone cannot reveal. All findings are delivered as publication-quality figures, interactive visual reports, and clear written interpretations designed for both scientific and non-scientific stakeholders — from your bench scientists to your leadership team.
Sitting on proprietary data that hasn't told its full story yet? We turn complex multi-omics datasets into findings your scientific and leadership teams can act on. Start with a free consultation →






















