Liquid Biopsy Intelligence: ctDNA in the Age of AI
1. What Is ctDNA? Biological Basis and Clinical Relevance
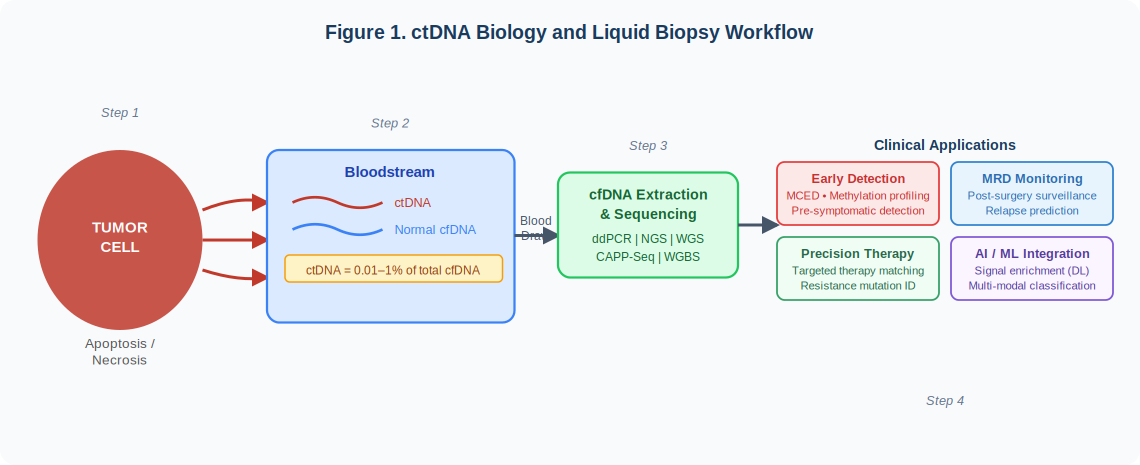
Circulating tumor DNA (ctDNA) refers to small fragments of cell-free DNA (cfDNA) shed by tumor cells into the bloodstream during apoptosis and necrosis. Unlike normal cfDNA derived from healthy tissues, ctDNA carries the somatic mutations, copy number alterations, and epigenetic modifications that define the originating tumor. These fragments are typically 160–200 base pairs in length and circulate at extremely low concentrations — often comprising as little as 0.01% to 1% of total plasma cfDNA.[1,2,20]
The clinical significance of ctDNA lies in its non-invasive accessibility: a simple blood draw can capture a whole-body molecular portrait of cancer biology that tissue biopsy cannot. Unlike single-site tissue sampling, which is spatially restricted and procedurally invasive, ctDNA provides a dynamic, real-time window into tumor evolution across all metastatic sites simultaneously. [20] This makes liquid biopsy — broadly defined as the analysis of tumor-derived materials in circulating blood — a paradigm-shifting tool for oncology.
Beyond mutations, an emerging sub-field called cfDNA fragmentomics leverages fragment size distributions, end-motif frequencies, and nucleosome positioning patterns as tissue-specific cancer signals. These features carry information about the chromatin architecture of the tumor cell of origin, enabling tissue-of-origin inference without relying solely on mutation data
Figure 1. ctDNA Biology and Liquid Biopsy Workflow. Tumor cells shed ctDNA into the bloodstream through apoptosis and necrosis. Following cfDNA extraction and sequencing, ctDNA analysis enables four clinical application domains: early cancer detection, MRD monitoring, precision therapy guidance, and AI/ML-assisted interpretation.
2. Clinical Applications of ctDNA Liquid Biopsy
ctDNA-based liquid biopsy has demonstrated utility across five major clinical domains:
2.1 Early Cancer Detection
Multi-cancer early detection (MCED) platforms analyze genome-wide cfDNA methylation profiles from a single blood sample to detect cancer signals before clinical presentation. Galleri (GRAIL), using deep learning on whole-genome bisulfite sequencing (WGBS) data, covers more than 50 cancer types. The NHS-Galleri trial enrolled 140,000 participants and demonstrated an overall sensitivity approaching 51%, with particularly strong performance for cancers lacking established screening tests. [12,15] PanSEER, a random forest classifier applied to cfDNA methylation, prospectively detected cancer up to four years before conventional diagnosis.
2.2 Molecular Residual Disease (MRD) Monitoring
Detection of residual ctDNA after curative-intent surgery predicts recurrence with remarkable precision. The landmark GALAXY study (n=2,240) established that ctDNA positivity post-resection predicted inferior disease-free survival (HR=11.99) and overall survival (HR=9.68) in colorectal cancer, [8] catalyzing regulatory interest in ctDNA MRD as a clinical trial endpoint.
2.3 Treatment Response Monitoring
Serial ctDNA quantification tracks tumor burden dynamics during therapy. Molecular responses — defined as significant ctDNA decline — frequently precede radiographic responses by weeks to months, enabling earlier clinical decision-making. [13] In multiple myeloma, ctDNA sequencing post-CAR T-cell therapy correlates strongly with bone marrow MRD status while simultaneously tracking CAR-specific cfDNA kinetics in a non-invasive manner.[17]
2.4 Genomic Profiling and Therapy Guidance
When tissue biopsy is infeasible, plasma ctDNA sequencing identifies actionable mutations (EGFR, ALK, KRAS, BRAF) for targeted therapy selection. The GOZILA study (n=4,037 GI cancer patients) validated ctDNA-guided matched therapy, achieving a 24% genomic match rate and significantly improved overall survival (HR=0.54) in matched versus unmatched patients.[14]
2.5 Resistance Mechanism Detection
Longitudinal ctDNA monitoring identifies emerging resistance mutations in real time, enabling adaptive therapy strategies before radiographic disease progression is detectable — a critical advantage in precision oncology where treatment timing directly affects outcomes.[13,17]
3. AI-Driven Data Analysis of ctDNA
The ctDNA signal space — spanning mutations, methylation, copy number variations, fragmentomics, end motifs, and nucleosome positioning — is extraordinarily information-rich but too high-dimensional for conventional statistical methods to fully exploit. Artificial intelligence and machine learning have emerged as essential tools for navigating this complexity.[2,5,19]
3.1 The Analysis Pipeline
A complete ctDNA AI pipeline proceeds through four stages:
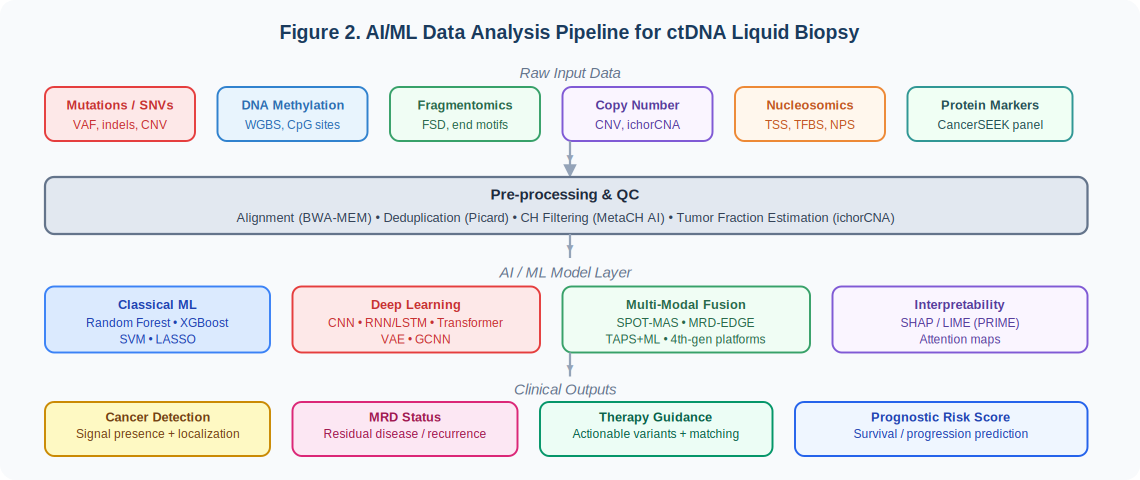
Raw data inputs: variant allele frequencies (VAF), CpG methylation matrices from WGBS, fragment size distributions, end-motif tetranucleotide frequencies, copy number profiles, and circulating protein biomarkers
Pre-processing and QC: read alignment (BWA-MEM), deduplication (Picard), somatic variant calling (MuTect2, Strelka2), tumor fraction estimation (ichorCNA), and AI-based clonal hematopoiesis (CH) filtering (MetaCH)[9]
AI/ML model layer: classical algorithms (Random Forest, XGBoost, SVM, LASSO) for structured tabular data; deep learning (CNN, RNN/LSTM, Transformer, VAE) for high-dimensional sequence and methylation features; multi-modal fusion architectures that jointly model all cfDNA feature types[2,5,19]
Clinical output interpretation: cancer signal presence and tissue-of-origin localization, MRD status, actionable variant identification, and prognostic risk scores with per-patient explainability via SHAP values.
Figure 2. AI/ML Data Analysis Pipeline for ctDNA Liquid Biopsy. Raw multi-modal cfDNA inputs (mutations, methylation, fragmentomics, CNV, nucleosomics, protein markers) flow through QC pre-processing into a tiered ML model layer, yielding four categories of clinical output: cancer detection, MRD status, therapy guidance, and prognostic risk scoring.
3.2 Landmark AI Tools
Several purpose-built AI frameworks have substantially advanced ctDNA analysis. MRD-EDGE (2024) integrates SNV and CNV signals from WGS using deep learning to achieve approximately a 300-fold improvement in signal-to-noise ratio over conventional WGS-based MRD detection, enabling tumor burden monitoring at ctDNA fractions below conventional analytical limits.[1]
SPOT-MAS (2023) simultaneously integrates methylomics, fragmentomics, CNV, and end-motif analysis within a single low-depth WGS workflow (~0.55x coverage), validated across 2,288 participants for multi-cancer detection. [3] MetaCH (2025) classifies cfDNA variants as CH-derived versus tumor-derived from plasma-only data — eliminating the need for matched white blood cell sequencing and substantially reducing false-positive rates. [9] PRIME (2026) is an interpretable multi-algorithm ML model that integrates ctDNA-MRD, mutation profiles, and clinicopathological variables to predict NSCLC progression risk across six independent cohorts, with SHAP-based per-patient explainability essential for clinical adoption.[10]
3.3 The Multi-Modal Frontier
The field is converging on fourth-generation liquid biopsy architectures that simultaneously analyze all cfDNA feature types within a single integrated ML framework. Pan-cancer multi-modal integration of mutations, CNV, methylation, fragmentomics, and end motifs achieves 72.4% sensitivity at 97% specificity across multiple cancer types — a performance frontier inaccessible to any single modality.[5] Future architectures will incorporate cfRNA, exosome proteomics, and metabolomics into unified AI frameworks, while foundation model approaches — analogous to those transforming single-cell genomics — promise to enable few-shot transfer to rare cancers from large-scale pretrained ctDNA representations.